March 2020, news was spreading of a rapidly evolving pandemic with potentially deadly concequences and societies were struggling to slow the tide.
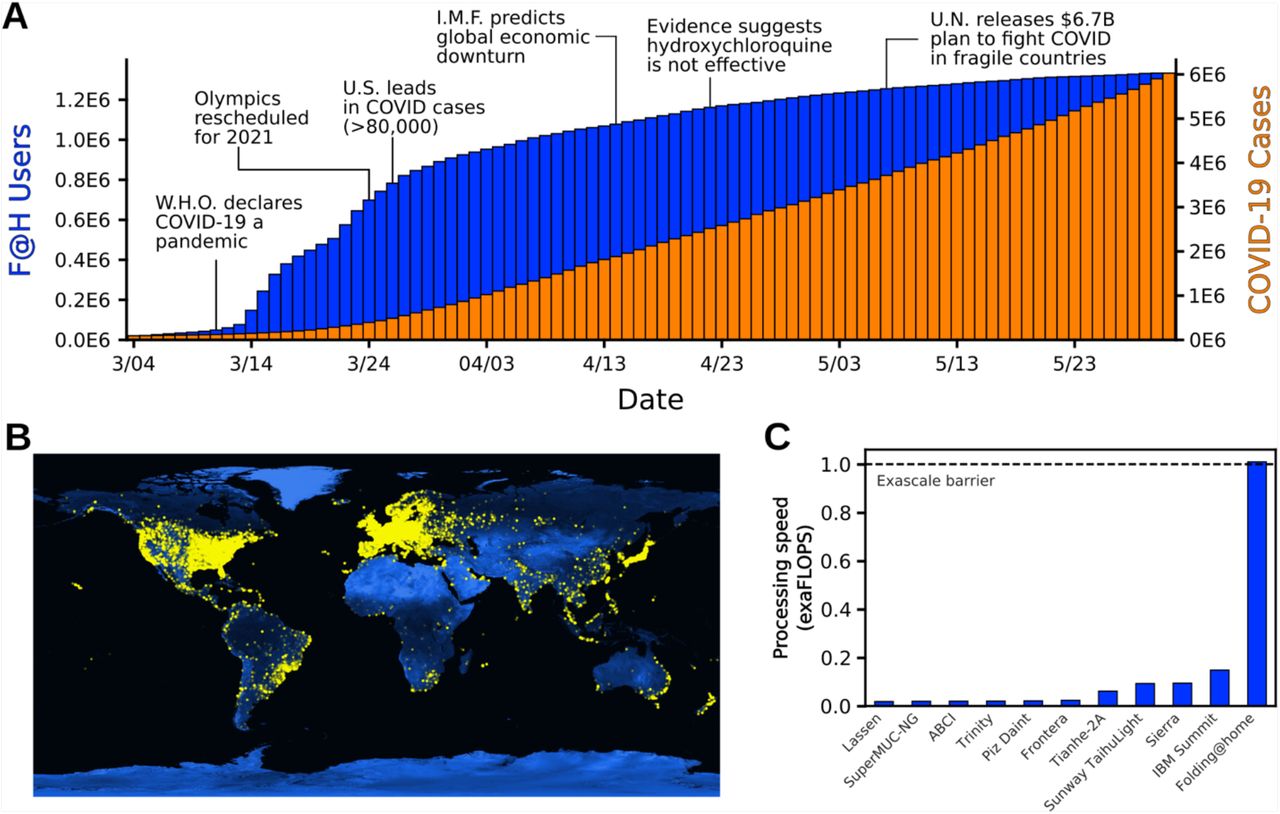
Less than a week before working remotely was made mandatory, in an effort to reduce human contact nationwide, I saw the news Folding@home update on SARS-CoV-2 (10 Mar 2020) the Folding@Home project was rapidly onboarding COVID-19/SARS-COV-2 protein folding projects to aid with early understanding of the virus in hopes of discovering ways to block the virus from interacting with human cells, preventing infection.
Every organisation on the planet was eager to do their part to blunt the pandemic and my role gave me access to large arrays of powerful compute - There wasn’t anything higher priority to be focusing on so I started to evaluate various research clusters suitability for this type of compute work, a few top considerations:
- Impact on critical production workloads
- Security considerations
- Rapid implementation
Any contributions we hoped to make couldn’t risk our critical production workloads, which given the pandemic were even more critical to the wellbeing of society than ever. This ruled out running these jobs on the production clusters, as there were too many unknowns related to the Folding@Home software and how it could affect the production environment.
We needed to avoid excessive security constraints, the Folding@Home software would not be vetted and would be downloading work units from the Internet and running automatically. This was quite concerning, as a malicious actor could infect the work units and potentially gain access to our clusters and by extension the wider internal network.
We had to move fast, it wasn’t long before events would grind everything to a halt, so if we wanted to maximize our contribution we had to preempt any concerns that higher-ups may have, and be in a position to launch as soon as approval was given. I was already testing the software and evaluating what type of environment it was capable of running in; whether containers would help with distribution, how much access to the Internet it needed, and whether it could integrate with our HPC batch schedulers.
The had stars aligned, an experimental research cluster used to evaluate exotic compute architectures was full of Intel Xeon Phi many-core CPUs, was not being used, was located in a heavily isolated network partition, could access the Internet, and was wholly controlled by us.
The HPC Team submitted our plan and rationale – The organisation had access to valuable spare compute that was in urgent need in the research community, the impact of our compute would be exponentially greater the earlier we contribute, and if they wanted a financial incentive: most, and soon to be all, employees were working from home, resulting in a much smaller electricity bill that could be diverted to this endeavour. Approval was given soon after.
The deployment was very basic, we installed the software to a shared location in the cluster and launched it using pdsh, since we had the system all to ourselves we didn’t have to worry around workload scheduling, and we pushed through necessary firewall changes in parallel.
Sooner than expected, the system filled up with work, and it looked like the Folding@Home software was happy to use every one of the 64 threads available on every compute node in the cluster!
TeamMO was founded by (Met Office) HPC Team and has earned 6,621,172,175 points by contributing 479,934 work units.

It’s hard to say what kind of impact this made, but I feel proud that in troubling circumstances we focused on what we could do to have a positive impact, and I’m thankful to my colleagues for supporting this effort so enthusiastically!
